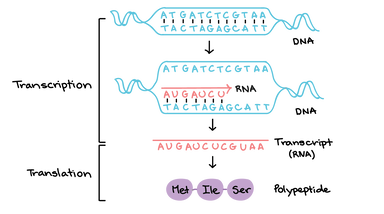This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is a transcriptome?
Transcriptomics is the study of the transcriptome- the entirety of RNA expressed in a certain cell under a specific set of conditions. RNA is expressed from genes in the genome through transcription (Figure 1). Analyzing the transcriptome allows scientists to conclude which genes are active in different processes, cell types, and environments. The two main assays for the transcriptome are RNA-sequencing and microarray analysis. [1]
What is RNA sequencing (RNA-seq)?
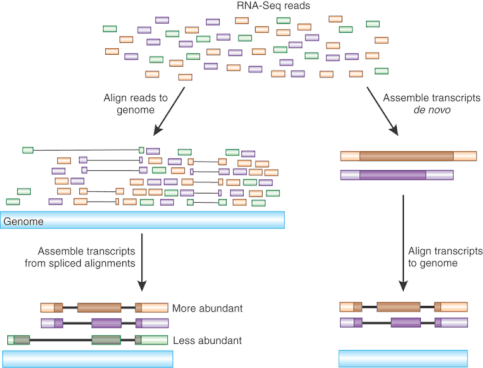
An RNA-seq experiment first creates cDNA (complementary DNA) based on RNA harvest from a cell. This DNA can then be sequenced to determine what RNAs are present in the cell. [2]
An RNA-seq experiment first creates cDNA (complementary DNA) based on RNA harvest from a cell. This DNA can then be sequenced to determine what RNAs are present in the cell. [2]
What is microarray analysis?
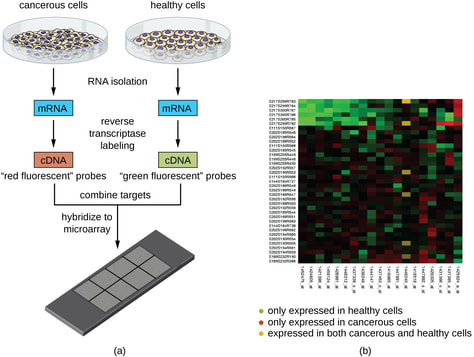
The DNA microarray is a tool used to study the extent to which certain genes are turned on or off in cells and tissues. RNA is isolated and the amount of transcripts from each gene is measured. [3]
The DNA microarray is a tool used to study the extent to which certain genes are turned on or off in cells and tissues. RNA is isolated and the amount of transcripts from each gene is measured. [3]
What is the expression pattern for LIPC?
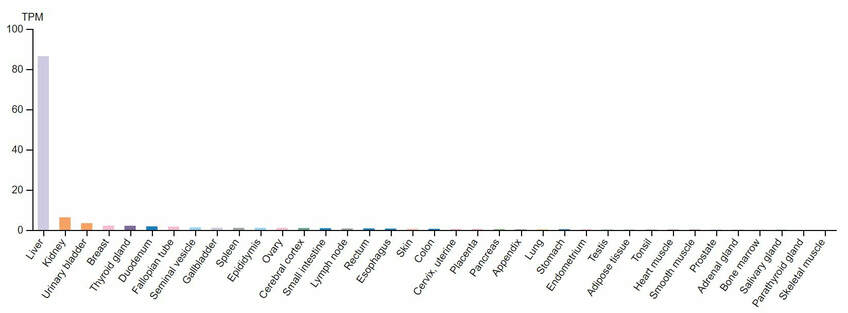
According to the Human Protein Atlas (HPA), LIPC is mainly expressed in the liver and adrenal glands. However there is some expression of LIPC in the breasts, thyroid, and female reproductive organs. Each of these places fits with the role of hepatic triacylglycerol lipase (HTGL), the protein expressed from LIPC, in cholesterol transport for hormone synthesis. A truncated form of LIPC is also expressed in the ovaries of mice, but human ovaries have not been checked for expression [4,5].
Conclusions
Transcriptomics is important for elucidating the role of different genes in certain biological processes and environments. While sequencing a genome gives us all the possible RNAs and proteins that can be created, only transcriptomics can tell us which ones are created and the situations in which they are used.
References
[1] Transcriptome Fact Sheet. (2015, August 27). Retrieved from https://www.genome.gov/13014330/
[2] RNA-Seq. (n.d.). Retrieved from https://www.sciencedirect.com/topics/neuroscience/rna-seq
[3] DNA Microarray Technology. (2015, August 27). Retrieved from https://www.genome.gov/10000533/dna-microarray-technology/
[4] Verhoeven, A.J., D. Carling, and H. Jansen, Hepatic lipase gene is transcribed in rat adrenals into a truncated mRNA. J Lipid Res, 1994. 35(6): p. 966-75.
[5] Verhoeven, A.J. and H. Jansen, Hepatic lipase mRNA is expressed in rat and human steroidogenic organs. Biochim Biophys Acta, 1994. 1211(1): p. 121-4.
Non-linked Figures:
Header
[2] RNA-Seq. (n.d.). Retrieved from https://www.sciencedirect.com/topics/neuroscience/rna-seq
[3] DNA Microarray Technology. (2015, August 27). Retrieved from https://www.genome.gov/10000533/dna-microarray-technology/
[4] Verhoeven, A.J., D. Carling, and H. Jansen, Hepatic lipase gene is transcribed in rat adrenals into a truncated mRNA. J Lipid Res, 1994. 35(6): p. 966-75.
[5] Verhoeven, A.J. and H. Jansen, Hepatic lipase mRNA is expressed in rat and human steroidogenic organs. Biochim Biophys Acta, 1994. 1211(1): p. 121-4.
Non-linked Figures:
Header