This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Gene Ontology (GO)?
The purpose of GO is to describe our knowledge of the biological domain of gene products. GO annotation consists of three different domains: molecular function, cellular component, and biological process.
|
Molecular Function:
Action completed by the gene of interest at the molecular level [1,2]. |
Cellular Component:
The location that contains the gene of interest [1,2]. |
Biological Process:
Overall accomplishment of a series of molecular functions [1,2]. |
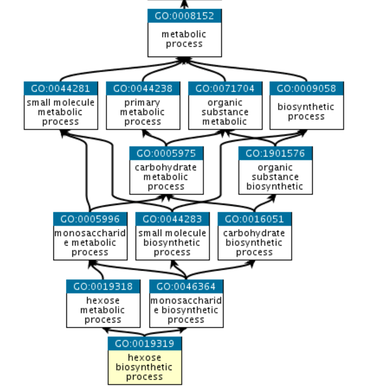
Gene ontology is somewhat hierarchical with more specific terms feeding into broader terms. This is illustrated with a graph similar to the one below.
Evidence for Assigning GO Annotations:
GO annotates terms with the evidence used to assign them. The evidence codes fall into six categories. Listed from most certain to least they are: experimental, phylogenetic, computational, author statements, curatorial statements, automatically generated annotations. Experimental codes indicate that this term has the support of a direct experimental assay, which is the most reliable form of evidence. A phylogenetic evidence code indicates that the term was assigned based on gene relationships, like homology. Computational evidence codes are based on in silico analysis of the gene/gene product sequence. Author statements codes state that the annotation was made based on a reference cited by an author in a paper and curatorial evidence is based on the GO curators judgement. Finally, electronic evidence is based on computer-generated evidence without human oversight.
GO annotates terms with the evidence used to assign them. The evidence codes fall into six categories. Listed from most certain to least they are: experimental, phylogenetic, computational, author statements, curatorial statements, automatically generated annotations. Experimental codes indicate that this term has the support of a direct experimental assay, which is the most reliable form of evidence. A phylogenetic evidence code indicates that the term was assigned based on gene relationships, like homology. Computational evidence codes are based on in silico analysis of the gene/gene product sequence. Author statements codes state that the annotation was made based on a reference cited by an author in a paper and curatorial evidence is based on the GO curators judgement. Finally, electronic evidence is based on computer-generated evidence without human oversight.
LIPC Gene Ontology
Here are the GO annotations for the gene product of LIPC, hepatic triacylglycerol lipase (HTGL). Each of the GO domains can be visualized in graphical form by clicking the titles. NaviGO is a website for visualizing GO term connections. The NaviGO links direct to the graphical view of LIPC GO annotations while the links in the description lead to the GO term mentioned [3].
|
Molecular Function:
HTGL is involved in phospholipase and triglyceride lipase activities. It binds to heparin, low-density lipoprotein particles, and apolipoproteins. HTGL is also involved in the regulation of lipoprotein lipase (LPL) activity. [4] |
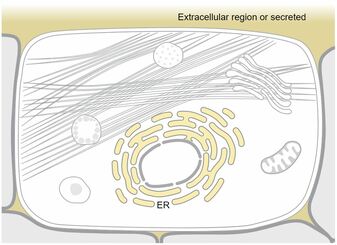
Cellular Component:
HTGL is found in the extracellular region and the endoplasmic reticulum lumen which are typical for secreted and membrane proteins. HTGL is also annotated as co-localizing with high-density lipoprotein particles which fits its role in cholesterol transport. [4] |
Biological Process:
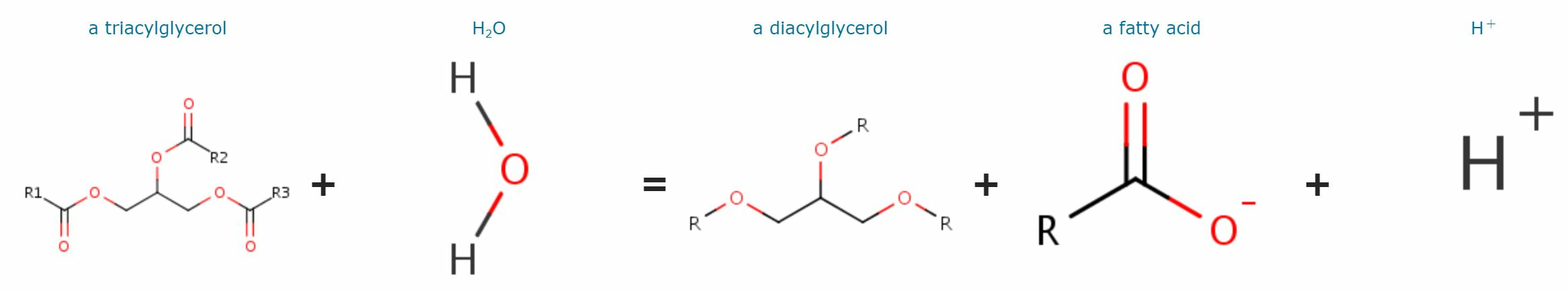
HTGL is involved in the cholesterol metabolic process, triglyceride catabolic process and cholesterol/triglyceride homeostasis. It also participates in the remodeling of very-low-density, low-density, intermediate-density, and high-density lipoproteins. HTGL plays a role in chylomicron remnant clearance. [4] |
Conclusion
Gene ontology is useful for understanding how hepatic triacylglycerol lipase (HTGL) functions in metabolism. Knowing the functions and interactions of HTGL can help scientists to design assays for the effects of different HTGL mutations. This will bring about better diagnostic testing for hepatic lipase deficiency.
References
[1] Ashburner et al. Gene ontology: tool for the unification of biology. Nat Genet. May 2000;25(1):25-9. doi:10.1038/75556
[2] The Gene Ontology Consortium. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. Jan 2019; 47(1):330-338. doi:10.1093/nar/gky1055
[3] Wei, Q., Khan, I. K., Ding, Z., Yerneni, S., & Kihara, D. (2017). NaviGO: Interactive tool for visualization and functional similarity and coherence analysis with gene ontology. BMC Bioinformatics,18(1). https://doi.org/10.1186/s12859-017-1600-5
[4] Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, Lewis S, AmiGO Hub, Web Presence Working Group. AmiGO: online access to ontology and annotation data. Bioinformatics. Jan 2009; 25(2):288-289. doi:10.1093/bioinformatics/btn615
Non-linked Figures:
Header: https://www.promega.com/products/protein-expression/
[2] The Gene Ontology Consortium. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. Jan 2019; 47(1):330-338. doi:10.1093/nar/gky1055
[3] Wei, Q., Khan, I. K., Ding, Z., Yerneni, S., & Kihara, D. (2017). NaviGO: Interactive tool for visualization and functional similarity and coherence analysis with gene ontology. BMC Bioinformatics,18(1). https://doi.org/10.1186/s12859-017-1600-5
[4] Carbon S, Ireland A, Mungall CJ, Shu S, Marshall B, Lewis S, AmiGO Hub, Web Presence Working Group. AmiGO: online access to ontology and annotation data. Bioinformatics. Jan 2009; 25(2):288-289. doi:10.1093/bioinformatics/btn615
Non-linked Figures:
Header: https://www.promega.com/products/protein-expression/