This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein domains?
Protein domains are units with distinct secondary structure that fold into a characteristic three-dimensional structure. They are responsible for a certain protein functions or interactions. Domains can be found in similar proteins with different functions. A protein may be assigned to a certain protein family based on the domains it contains. [1,2]
How do scientists identify protein domains?
First scientists must build an alignment of sequences from related proteins. From there, they can identify pieces of the sequence that are conserved across the protein family. Once a pattern emerges, tests can be done to investigate its functional and structural relevance. This information is used to define parameters for the domain. Once a complete definition is generated, the domain is added to protein databases and newly discovered proteins can be compared to it to see if they contain the domain. Many databases for protein domain analysis exist. Some of the more commonly used databases are listed below. [3]
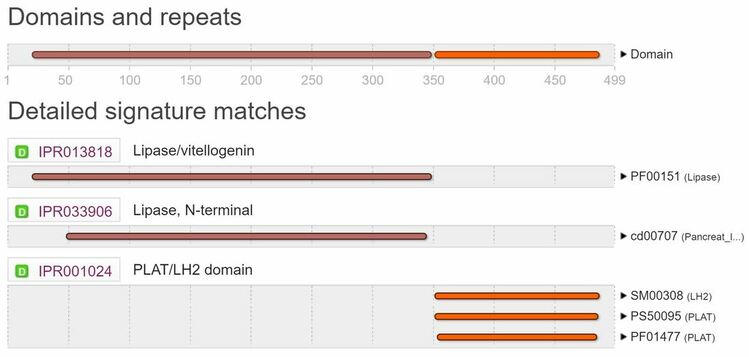
What domains does Hepatic Triacylglycerol Lipase (HTGL) have?
|
Lipase Domain:
Triglyceride lipases are enzymes that hydrolyze ester linkages of triglycerides [4]. The most conserved region in all these proteins is centered around a serine residue (pink flag in Figure 2) which participates in a charge relay system along with histidine and aspartic acid residues (red flags in Figure 2) [5]. |
PLAT/LH2 Domain:
This is a domain found in a variety of membrane or lipid associated proteins. It has two names: PLAT (Polycystin-1, Lipoxygenase, Alpha-Toxin) domain or LH2 (Lipoxygenase homology) domain. The structure of this domain is a beta-sandwich composed of two sheets of four strands each [6,7,8]. Most of the highly conserved residues are buried within the protein except for a surface lysine or arginine which may be a conserved ligand-binding site. Overall, this domain may mediate membrane attachment via other protein binding partners. |
Hepatic Triacylglycerol Lipase Domain Conservation
Both the lipase and PLAT/LH2 domains in HTGL are conserved across its orthologs. The architecture of these domains differs slightly in the paralogs, but the lipase domain is conserved in all proteins. The orthologs all contain both domains, but with different positions and protein lengths.
Both the lipase and PLAT/LH2 domains in HTGL are conserved across its orthologs. The architecture of these domains differs slightly in the paralogs, but the lipase domain is conserved in all proteins. The orthologs all contain both domains, but with different positions and protein lengths.
Conclusions
Hepatic triacylglycerol lipase (HTGL) consists of two main domains, the lipase domain and the PLAT/LH2 domain. The lipase domain is responsible for the hydrolysis of triglycerides which is the main function of the enzyme. The PLAT/LH2 domain may allow HTGL to bind to other proteins or assist in membrane binding. Due to the abundance of genomes available after modern sequencing, domain analysis allows scientists to rapidly classify proteins based on structures and functions of known proteins.
References
[1] Family- and domain-based protein classification. (2016, June 08). Retrieved from https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/family-and-domain-based-protei
[2] European Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. (n.d.). European Bioinformatics Institute. Retrieved from https://www.uniprot.org/help/domain
[3] Mulder, N. J., & Apweiler, R. (2001). Tools and resources for identifying protein families, domains and motifs. Genome biology, 3(1), REVIEWS2001. (link)
[4] Persson, B., Bengtsson-Olivecrona, G., Enerback, S., Olivecrona, T., & Jornvall, H. (1989). Structural features of lipoprotein lipase. Lipase family relationships, binding interactions, non-equivalence of lipase cofactors, vitellogenin similarities and functional subdivision of lipoprotein lipase. European Journal of Biochemistry,179(1), 39-45. doi:10.1111/j.1432-1033.1989.tb14518.x
[5] Blow, D. (1990). Enzymology. More of the catalytic triad. Nature,343(6260), 694-695. doi:10.1038/343694a0
[6] Bateman, A., & Sandford, R. (1999). The PLAT domain: A new piece in the PKD1 puzzle. Current Biology,9(16). doi:10.1016/s0960-9822(99)80380-7
[7] Delrieu, I., Waller, C. C., Mota, M. M., Grainger, M., Langhorne, J., & Holder, A. A. (2002). PSLAP, a protein with multiple adhesive motifs, is expressed in Plasmodium falciparum gametocytes. Molecular and Biochemical Parasitology,121(1), 11-20. doi:10.1016/s0166-6851(02)00016-6
[8] Tomchick, D. R., Phan, P., Cymborowski, M., Minor, W., & Holman, T. R. (2001). Structural and Functional Characterization of Second-Coordination Sphere Mutants of Soybean Lipoxygenase-1†. Biochemistry,40(25), 7509-7517. doi:10.1021/bi002893d
Non-linked Figures:
Header: https://www.rcsb.org/structure/1hpl
[2] European Bioinformatics InstituteProtein Information ResourceSIB Swiss Institute of Bioinformatics. (n.d.). European Bioinformatics Institute. Retrieved from https://www.uniprot.org/help/domain
[3] Mulder, N. J., & Apweiler, R. (2001). Tools and resources for identifying protein families, domains and motifs. Genome biology, 3(1), REVIEWS2001. (link)
[4] Persson, B., Bengtsson-Olivecrona, G., Enerback, S., Olivecrona, T., & Jornvall, H. (1989). Structural features of lipoprotein lipase. Lipase family relationships, binding interactions, non-equivalence of lipase cofactors, vitellogenin similarities and functional subdivision of lipoprotein lipase. European Journal of Biochemistry,179(1), 39-45. doi:10.1111/j.1432-1033.1989.tb14518.x
[5] Blow, D. (1990). Enzymology. More of the catalytic triad. Nature,343(6260), 694-695. doi:10.1038/343694a0
[6] Bateman, A., & Sandford, R. (1999). The PLAT domain: A new piece in the PKD1 puzzle. Current Biology,9(16). doi:10.1016/s0960-9822(99)80380-7
[7] Delrieu, I., Waller, C. C., Mota, M. M., Grainger, M., Langhorne, J., & Holder, A. A. (2002). PSLAP, a protein with multiple adhesive motifs, is expressed in Plasmodium falciparum gametocytes. Molecular and Biochemical Parasitology,121(1), 11-20. doi:10.1016/s0166-6851(02)00016-6
[8] Tomchick, D. R., Phan, P., Cymborowski, M., Minor, W., & Holman, T. R. (2001). Structural and Functional Characterization of Second-Coordination Sphere Mutants of Soybean Lipoxygenase-1†. Biochemistry,40(25), 7509-7517. doi:10.1021/bi002893d
Non-linked Figures:
Header: https://www.rcsb.org/structure/1hpl